Recently Published

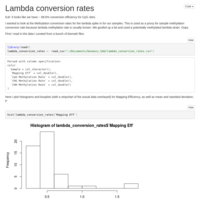
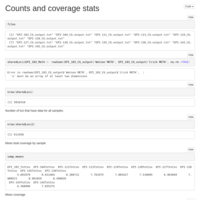
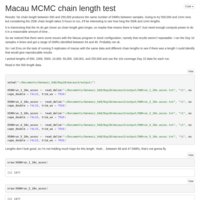
5 Sample 10x Coverage C virginica MACAU run
Re-tried MACAU kicking out the one poor quality sample. Got a few more meaningful loci to look at, but all had Beta values that were indistinguishable from 0.
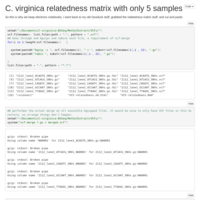
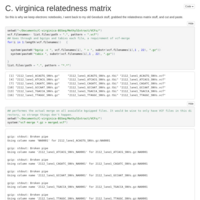
5 sample relatedness matrix
Just like the 6 sample version, only with 5!
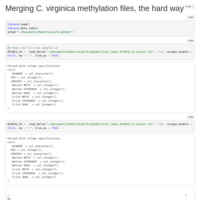
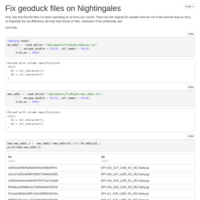
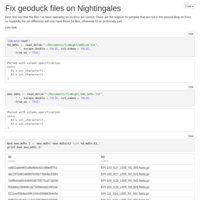
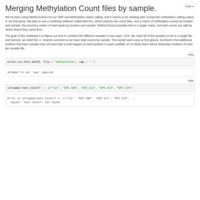
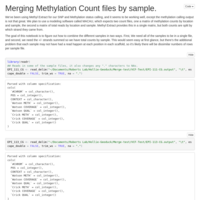
Merging C. virginica methylation count files
files were too big/disparate for R to handle, sqlshare too big of a pain, so I did it the hard way!
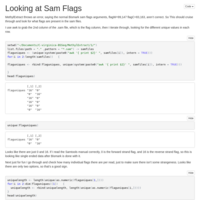
Finding the right flags for the C virginica bismark files
I'm sure there's an easier way to do this, but it seemed to work.

Methylkit success!
Just needed to change the flagW and flagC arguments to 0 and 16 respectively.
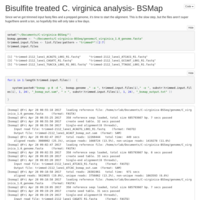
C. virginica methylation analysis, pt 6
BSMap comparison for trimmomatic trimmed files
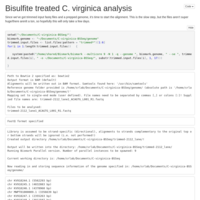
C. virginica methylation analysis, pt 5
Re-ran Bismark with re-trimmed files. No measurable change in mapping rates.
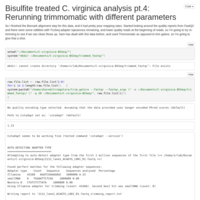
C virginica methylation analysis pt.4
Re-ran trimming step using Trimmomatic and TruSeq adapter sequences specified.
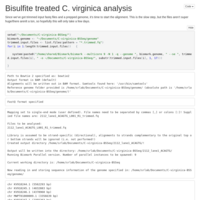
C virginica methylation analysis pt.3
Bismark run on C. virginica oil exposure samples. Super low mapping rate (sub 10%) which is strange...
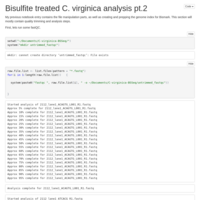
C. virginica methylation analysis, pt 2
Fastqc, trimming, and re-fastqcing

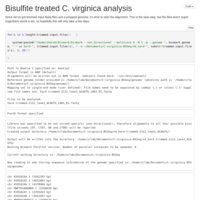
C. virginica methylation analysis, pt 1
Moving and checking files, prepping the genome.
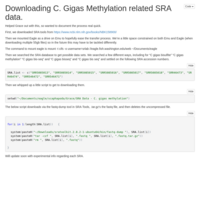
Updated SRA Download script
This time it works!
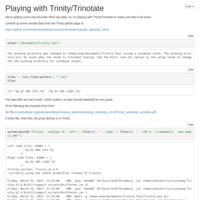
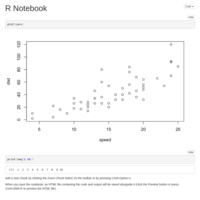
Test for Marcos
Testing upload for R Notebook

Experimenting with MethPipe
or: Why scientists shouldn't be allowed to name things.

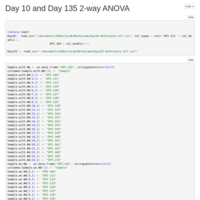
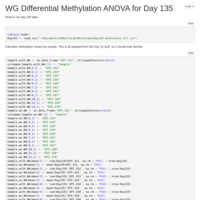
Day 10 Whole Genome Methylation ANOVA
Tried an ANOVA on mean methylation for Day 10 samples by treatment. Nothing really interesting came of it. On to day 135!
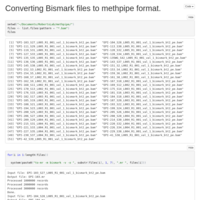
MethPipe Prep
Forgot to upload this. Super time/space intensive but its nearly done after a lot of hard drive shuffling!
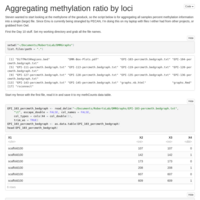
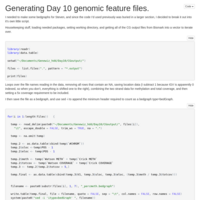
Methylome data file
Making a file of percent methylation to start looking at the geoduck methylome.

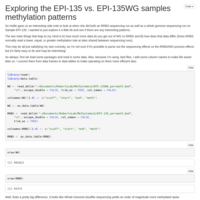
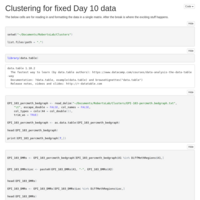
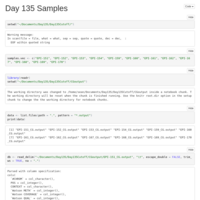
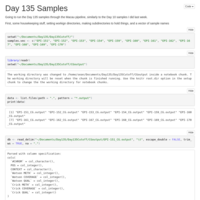
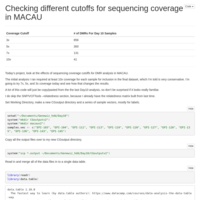
Day 10 and Day 135 analysis
Looked at the combined Day 10 and Day 135 samples, with the inclusion of a time covariate.
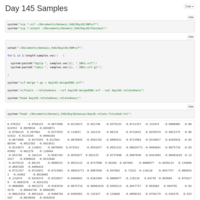
Day 145
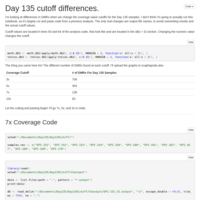
These are the day 145 runs, with the corrected methylation cutoff logic. Macau was run outside of R-studio, to facilitate running multiple instances concurrently.
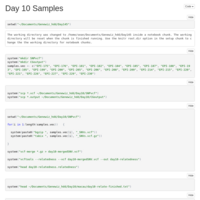
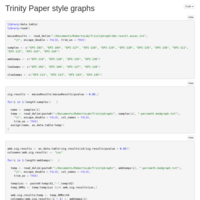
Day 10, done correctly
So I noticed that I was inadvertently trimming all methylation counts less than 10, as well as the total counts. I should not have done that. The good news is this nets us 267 DMRs as opposed to 41! The bad news, for some reason it makes MACAU run glacially slow.

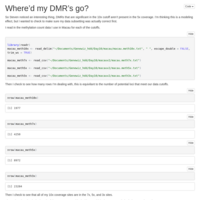
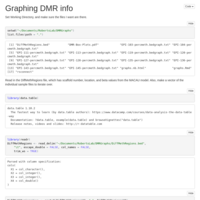
vcf-merge and vcftools --relatedness
Pseudocode description on how to combine multiple SNP files in to a single merged file, and then generate a relatedness matrix.

Methyl Extract, one at a time
Running Methyl Extract on a single sample .sam file at a time.

TA protocol
Rough protocol outline after meeting with Micah at DNR in Olympia.
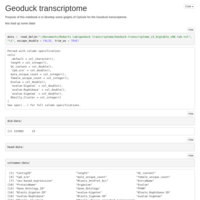
CpG O/E Forest Plot
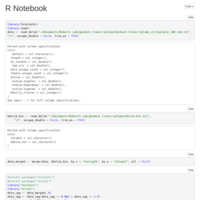
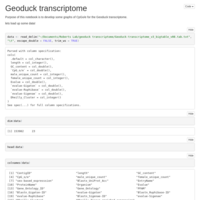
CpG O/E plot for Geoduck transcriptome, try 2.
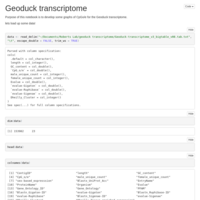
CpG O/E
Forest Plot code for Geoduck transcriptome methylation

MethylKit output, test run
Testing R notebook functionality with some Methylkit output from Steven's Oly samples.